Introduction
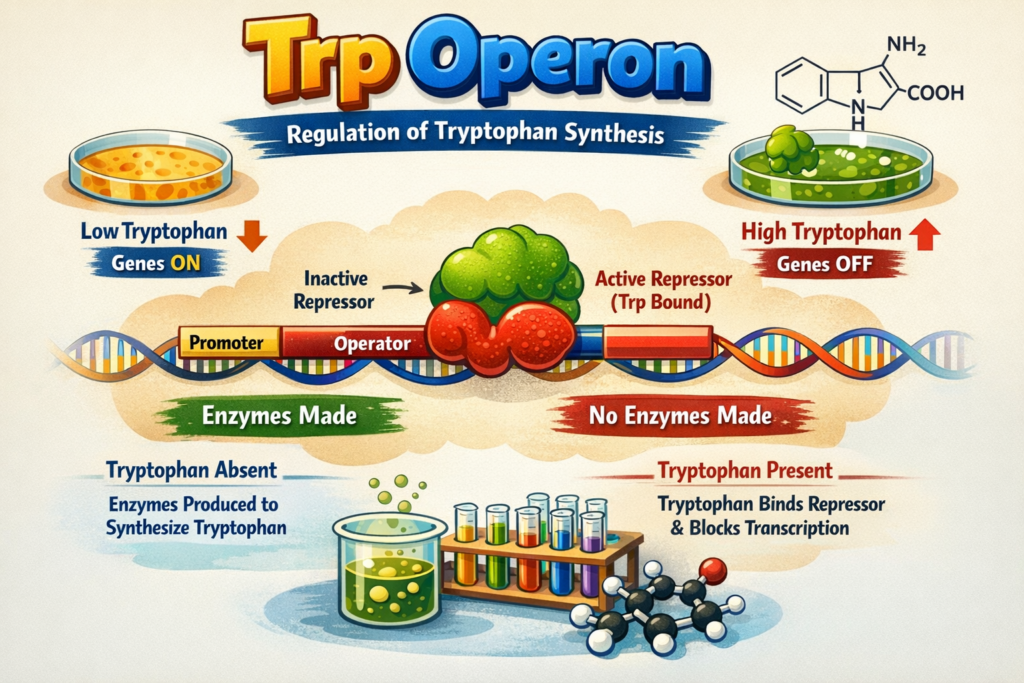
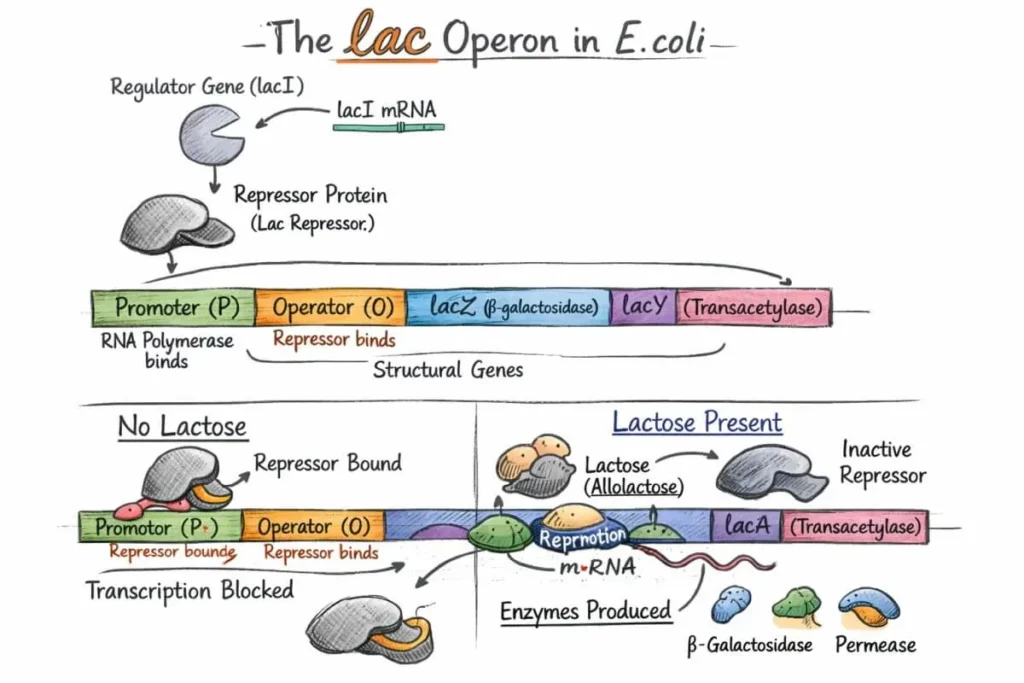
Genetic methylation test also known as DNA methylation test.Epigenetics looks at how certain changes can switch genes on or off without disturbing DNA sequence itself. One of the main ways this happens in mammals is through DNA methylation. Basically, a methyl group attaches to the fifth carbon of a cytosine, but only if that cytosine comes right before a guanine , scientists call it a CpG site. These CpG sites tend to gather in stretches of DNA that are rich in CG pairs, known as CpG islands. CpG island mostly presesnt on upstream region of the gene, like promoters, but sometimes they pop up in places like introns or exons too.
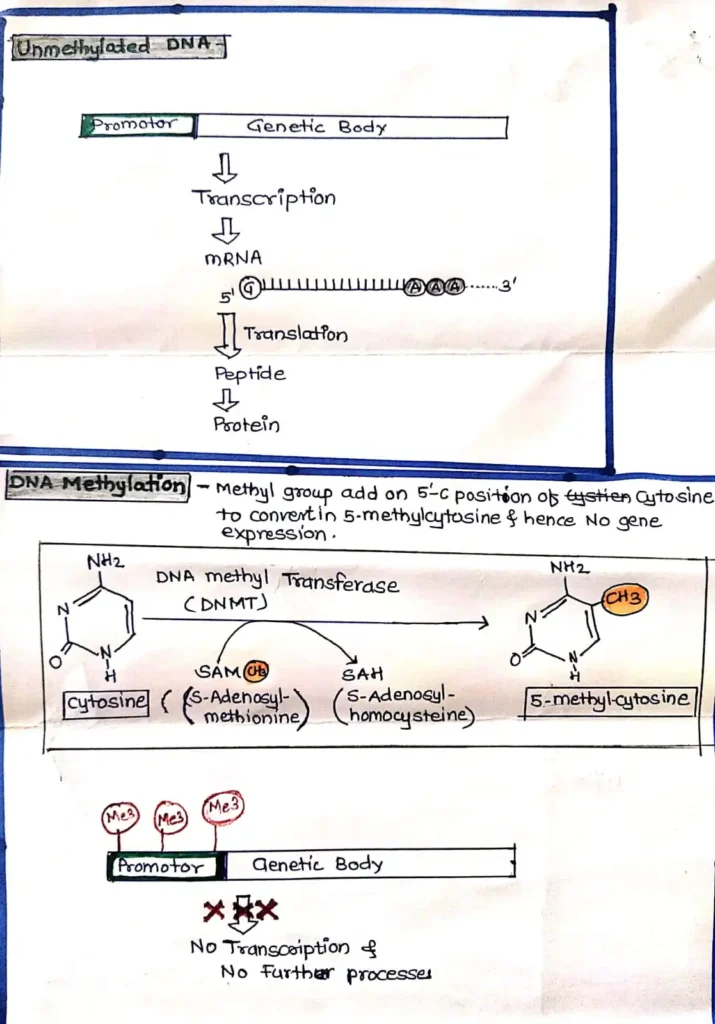
Researchers have also noticed methylation happens at “CGI shores,” which are areas with fewer CpG sites sitting right next to CpG islands. DNA methylation or Genetic methylation test does a lot of jobs in normal cells. It shuts down transposable elements, keeps viral sequences quiet, helps maintain the structure of chromosomes, inactivates one of the X chromosomes in females, and generally keeps a bunch of genes turned off when they’re not needed. The cool part is Cells copy these methylation patterns very precisely every time they divide.
As people age or develop certain diseases, these patterns can shift. Changes in DNA methylation stand out in a lot of complex diseases—heart disease, diabetes, neurological disorders—but researchers have dug deepest in cancer. Because of this, many teams have worked to pinpoint unique DNA methylation or Genetic methylation test “signatures” tied to how these diseases develop. These signatures actually help. They can serve as biomarkers for diagnosis, prognosis, tracking disease, or even predicting how someone will respond to treatment.
In short,Genetic Methylation is the process of adding a small chemical group called “methyl (–CH₃)” to DNA.This process does not change the DNA codes (genes), but it determines which genes will remain ON and which will remain OFF.
How to Choose Right method for Genetic Methylation Test
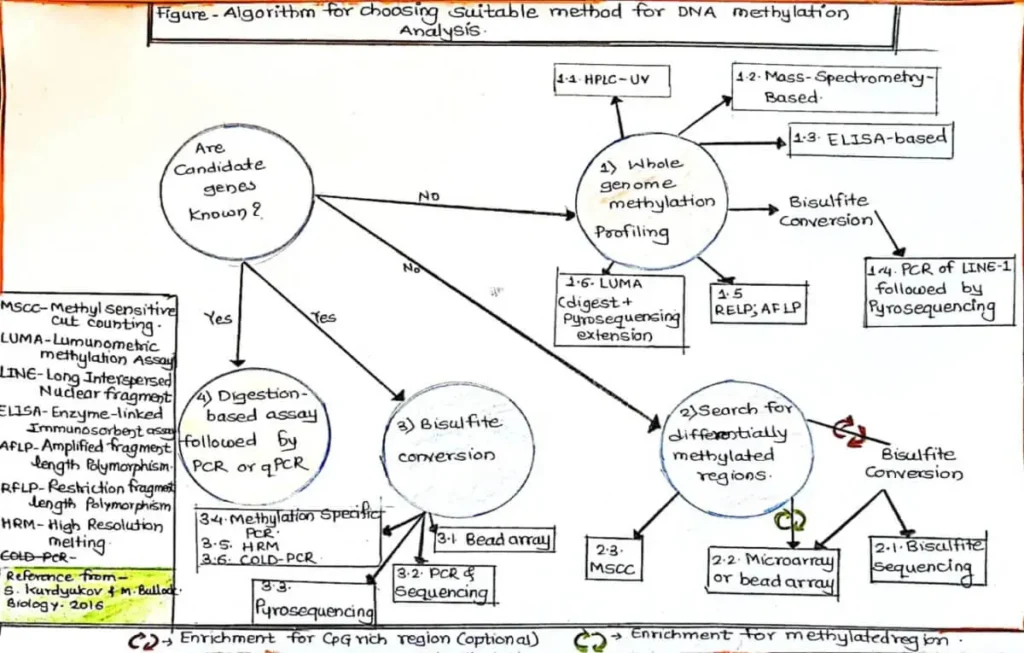
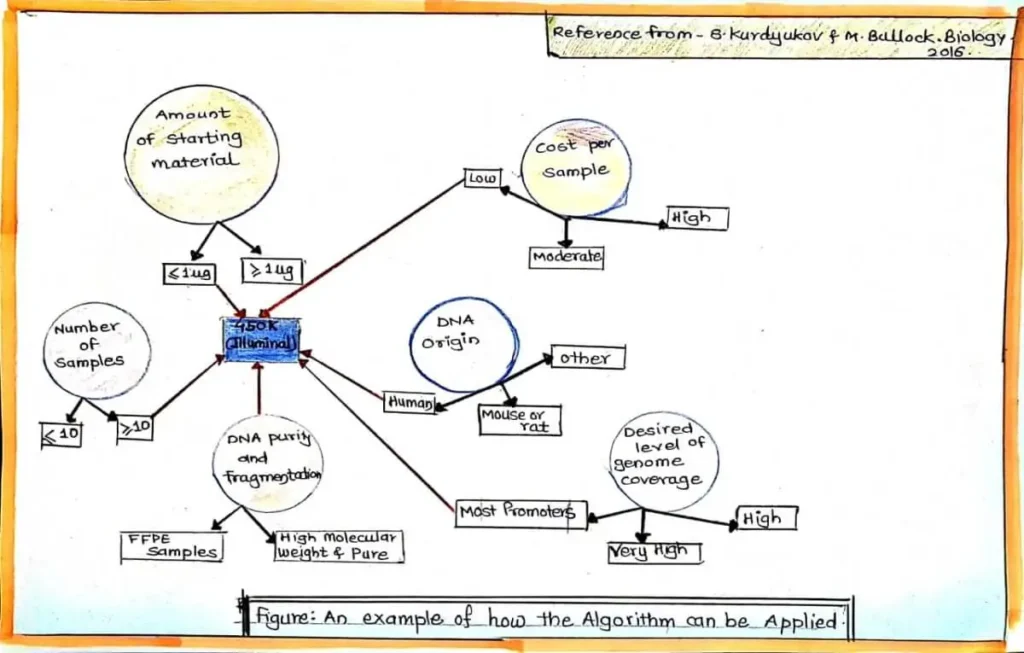
Main Strategies or Methods of Genetic Methylation Test
As the methylation of DNA data is diminished during PCR amplification, the most methods depend on methylation-based preparation of DNA before amplification. Genetic methylation test can be divided into three primary groups: A) Restriction enzyme (RE) digestion, B) Affinity-Based Analysis, and C) Bisulfite treatment. The union of these three methods involving sequencing and microarray-based technologies has led to an extensive array of techniques for global analysis of DNA methylation i.e Genetic Methylation. Sometime this three strategies mixed together for genetic methylation test like Combined bisulfite restriction analysis (COBRA) and Combination of methylated-DNA precipitation and methylation-sensitive restriction enzymes (COMPARE-MS).
A) Restriction enzyme (RE) digestion
It is one of the strategies of Genetic methylation test. Restriction Enzyme based methods utilize the property of Methylation Sensitive enzyme(Only digest unmethylated DNA) and Methylation Dependent Enzyme (Only cut Methylated DNA). These enzymes help identify methylated or unmethylated DNA sequences and show the level of Genetic methylation test.
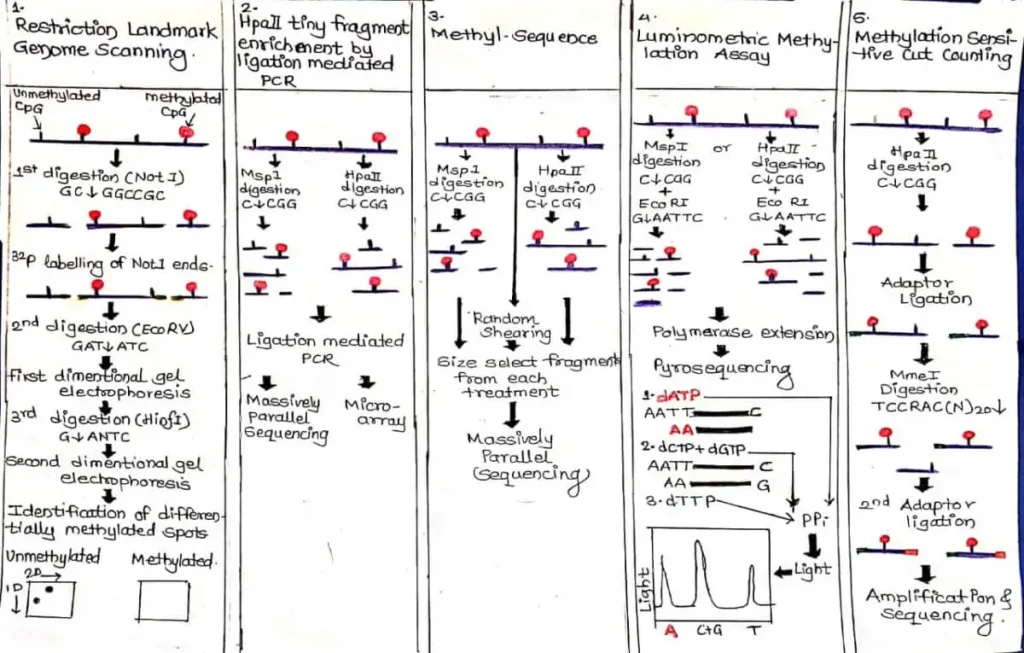
Techniques used in Restriction Enzyme (RE) Digestion-
1) Restriction Landmark Genome Sequencing(RLGS).
Principle
RLGS was the first dependable RE-based method for analyzing global Genetic methylation test. As it involves radioactive materials and gel electrophoresis the use of RLGS is declining.Refer FIGURE A.
Process
In this method ,Genomic DNA is digested or cleaved with methylation-sensitive restriction Enzyme,Lebelled them, and to visualize thousand of spot ; seperated by using gel eletrophoresis.Detection is based on ,methylated sites prevent digestion and hence not produce radioactive spot on gel,indicating hypermethylation.
Advantage:
- without requiring prior sequencing knowledge ,provide a quantitative reproduible map of methylation.
- Reproducibility and Accuracy – Give dependable profiles that show the details of CpG islands.
- Comprensive Coverage – Identifies Methylation changes in CpG island,promotor, target gene .
- Cost-effective – Less expensive than other method.
- low input -In case of low amount of DNA,it shows well function with accuracy.
- Easy to perform.
Disadvantage Or Limitation:
- Technically complex – RLGS is a technically complex procedure and requires specialized equipment, expertise, and experience.
- High-quality DNA required – Large amounts of high-quality DNA are required. Low or poor quality DNA does not yield accurate results.
- Low sensitivity – Minor methylation changes or small changes cannot be easily detected by RLGS.
- Limited to restriction enzyme sites – RLGS only analyzes the regions where restriction enzymes cut; the entire genome is not fully represented.
- Data analysis is difficult and time-consuming – It is difficult to understand and analyze the spot patterns that form.
- Not suitable for high-throughput – RLGS is not effective for testing a large number of samples at once.
- Less accurate than modern methods – RLGS is less accurate and less informative than modern techniques such as bisulfite sequencing or Next Generation Sequencing (NGS).
Application :
- Genome-wide DNA methylation pattern studies– RLGS is used to identify where methylation occurs across the entire genome.
- Uses in cancer research – RLGS is used to detect differences in DNA methylation between normal cells and cancer cells.
- Identifying tumor suppressor genes – This method is useful for detecting tumor suppressor genes that have been silenced due to methylation.
- Study of Epigenetic Changes – RLGS is used to understand epigenetic changes (changes that occur without changing the DNA sequence) that affect gene expression.
- Comparison of normal and disease states – RLGS is used to compare methylation patterns between healthy (normal) and diseased (disease) cells.
- Uses in developmental biology – Used to study methylation changes that occur during growth, development, and differentiation.
- Biomarkers identification – RLGS is useful for identifying certain methylation patterns as disease biomarkers.
- Gene regulation studies – Methylation analysis is used to understand which genes are turned ON or OFF.
2) HpaII Tiny Freagments Enrichment by Ligation Mediated PCR (HELP).
In HpaII Tiny Freagments Enrichment by Ligation Mediated PCR ,high molecular weight genomic DNA Digestion with Methylation Sensitive HpaII .Refer FIGURE A.
Principle–
(HpaII Tiny Fragments Enrichment by Ligation-Mediated PCR) is a method for DNA methylation analysis.The basic principle of this technique is:
- The HpaII restriction enzyme recognizes the CCGG sequence in DNA.
- If there is no methylation at that CCGG sequence, HpaII cuts the DNA there.
- If there is methylation at that site, HpaII cannot cut the DNA.
That is, DNA methylation patterns are identified by comparing DNA fragments cut (unmethylated) with uncut (methylated) fragments by HpaII.
The principle of this method is based on the difference in the behavior of two restriction enzymes, HpaII and MspI.
- HpaII is a methylation-sensitive enzyme. (If the CCGG sequence is methylated, HpaII does not cut and If it is unmethylated, HpaII cuts.)
- MspI is a methylation-insensitive enzyme.(It cuts whether the CCGG sequence is methylated or not.)
- By comparing the DNA fragments produced by both enzymes, it is determined which parts are methylated and which are not.
Process –
i)Genomic DNA Extraction – High quality genomic DNA is isolated from cells/tissue.
ii)Restriction Enzyme Digestion – DNA is split into two parts:HpaII digestion(HpaII cuts only at unmethylated CCGG sites.)
MspI digestion(MspI cuts at all CCGG sites (control/reference).
iii)Tiny Fragments Enrichment – Small DNA fragments (tiny fragments) formed after HpaII digestion are selected.These fragments often represent CpG-rich and unmethylated regions.
iv) Adapter Ligation – Synthetic adapters are added to the ends of selected DNA fragments.These adapters are required for subsequent PCR amplification.
v)Ligation-Mediated PCR – Adapter-ligated fragments are amplified by PCR.This increases the amount of DNA and facilitates further analysis.
vi)Comparative Analysis – HpaII-derived and MspI-derived fragments are compared.Regions in which HpaII signal is low/absent but MspI signal is present i.e methylated region and Regions in which both signals are present i.e unmethylated region.
Advantage:
- Accurate information about Genetic methylation test can be obtained through the HELP technique. The methylation/unmethylation at CpG sites in the genome can be accurately identified.
- High-throughput methods can study the methylation patterns of thousands of genomic regions at once, making them useful for large-scale studies.
- Quantitative Analysis Possible – The HELP method not only indicates the presence/absence but also the degree of methylation.
- Highly sensitive to unmethylated CpG regions – The HpaII enzyme only cuts unmethylated CpG sites, making the analysis of unmethylated regions more accurate.
- Useful for Cancer and Epigenetics Research: HELP is very effective in detecting abnormal DNA methylation changes that occur in cancer.
- The specificity is higher because the restriction enzyme–based method is more specific. Comparison of HpaII and MspI enzymes clearly demonstrates methylation-specific differences.
- Genome-wide methylation profiling is possible. It has the ability to study methylation patterns across the entire genome.
- Can be used with microarray or sequencing – HELP data can be combined with microarray or next-generation sequencing for more in-depth analysis.
Disadvantage :
- Enzyme-based restriction -The HELP technique relies on the restriction enzyme HpaII. HpaII only works on a specific sequence, CCGG, and therefore does not provide methylation information for other parts of the genome.
- Genome Coverage Remains Incomplete -Not all CpG sites in the entire genome are included in the CCGG. Therefore, the methylation profile obtained by the HELP method may be limited and incomplete.
- The Problem of PCR Bias -During ligation-mediated PCR, some fragments are amplified more and some are not. This can lead to bias in the results and the true methylation levels are not accurately reflected.
- Low quantitative accuracy -This method is not completely quantitative. That is, it is difficult to accurately determine the percentage of methylation; most often a relative comparison can be made.
- High-quality DNA required -HELP requires good quality, intact genomic DNA. Using degraded or low-quality DNA will not yield reliable results.
- Technical Complexity -This method involves many steps (digestion, ligation, PCR). Therefore, the possibility of error is high and the experiment is relatively difficult to perform.
- Problem with Repeat Regions – Repetitive sequences in the genome do not map properly. Therefore, methylation information of such regions may be missed.
- Less Sensitive Than Next-Generation Methods – The HELP method is less sensitive and less comprehensive than bisulfite sequencing or NGS-based methylation techniques.
- Time-consuming process – The experiment takes a long time to complete because of the many steps.
- Interpretation is difficult – Analysis and biological interpretation of the data obtained are difficult without expert knowledge.
3) Methyl seq.
In Methyl-Seq, DNA is either digested with MspI, HpaII, or randomly sheared. The digestion products are size fractioned and the selected fragments are sequenced.Refer FIGURE A.
Principle
Restriction enzyme–based Methyl-Seq is a method used to identify DNA methylation, which is based on the difference in cutting (digestion) between methylation-sensitive restriction enzymes (e.g. HpaII) and methylation-insensitive enzymes (e.g. MspI). Methylated vs unmethylated regions are identified by NGS(Next-Generation Sequencing) sequencing of the resulting fragments.
Process
i) Genomic DNA Extraction – Pure and high-quality DNA is isolated from cells or tissues.
ii) DNA Fragmentation – DNA is broken into small pieces (fragments) (sonication/enzymatic).
iii) Fragment Selection / Size Selection – Size-selected fragments are selected from the prepared fragments (more CpG-rich regions are obtained).
iv) Adapter Ligation – NGS adapters are attached to the ends of selected fragments.
v) PCR Amplification – A library is created by PCR amplification of adapter-ligated fragments.
vi) Next-Generation Sequencing – The prepared library is sequenced on an NGS platform (e.g. Illumina).
vii) Data Analysis – Sequencing reads are aligned to the reference genome. Methylated and unmethylated regions are determined by comparing read counts from sensitive vs insensitive digestion.
Advantage
- No bisulfite treatment required – Less DNA damage, relatively gentle process.
- Cost-effective – Cheaper than whole-genome bisulfite sequencing.
- NGS-based high throughput – Multiple samples can be analyzed simultaneously.
- Focus on CpG-rich regions – Useful for promoters and regulatory regions.
- Relative quantitative analysis – Easy comparison of methylation patterns.
Disadvantage
- Restriction site-dependent coverage – Only provides information on the sequences where the enzyme cuts; not complete genome coverage.
- No single-base resolution – Not as accurate as Bisulfite-based Methyl-Seq.
- Enzyme bias – Some genomic regions may be over/underrepresented.
- Difficulty with repeat regions – Mapping difficult; some parts are missed.
- High-quality DNA required – Poor DNA quality affects digestion and sequencing results.
4) Luminometric Methylation Assay.(LUMA)
In LUMA, DNA is digested with HpaII or MspI followed by digestion with EcoRI, bioluminometric polymerase extension, and pyrosequencing.Refer FIGURE A.
Principle
Luminometric Methylation Assay (LUMA) is a method used to measure DNA methylation, which is based on the difference in digestion between methylation-sensitive restriction enzyme (HpaII) and methylation-insensitive restriction enzyme (MspI). The amount of cuttings produced after HpaII and MspI digestion is measured by Pyrosequencing-based luminometric detection. From this, the genome-wide methylation level (%) is determined.
Process
i) Genomic DNA Extraction – Pure genomic DNA is isolated from cells or tissues.
ii) Restriction Enzyme Digestion – DNA is digested in two separate reactions:(HpaII + EcoRI digestion and MspI + EcoRI digestion) (EcoRI is used as an internal control).
iii) Pyrosequencing Reaction – Nucleotides are sequentially added to digested DNA. Light (luminescence) is produced as each nucleotide is incorporated.
iv) Luminometric Signal Detection – Light intensity is measured. The ratio of the HpaII digestion signal to the MspI digestion signal is calculated.
v) Methylation Calculation – Total DNA methylation (%) is calculated using the HpaII / MspI ratio.
Advantage
- Fast and high-throughput method – Many samples can be analyzed in a short time.
- Genome-wide methylation estimate – The average methylation level of the entire genome is known.
- No bisulfite treatment is required- DNA damage is reduced.
- Quantitative result – Methylation is obtained in percentage.
- Low DNA requirement – Even a small amount of DNA is sufficient.
Disadvantage
- No single-site or gene-specific information is available – Only global methylation level is available.
- Restriction site dependent method – Analysis is based on CCGG sites only.
- Low resolution – No CpG-specific or single-base resolution.
- Repeat regions and CpG islands cannot be distinguished – Functional interpretation is limited.
- Pyrosequencing instrument required – May not be available in all laboratories
5) Methylation Sensitive Cut Counting( MSCC).
Methyl-Sensitive Cut Counting (MSCC) is a restriction enzyme-based sequencing method for Genetic methylation test.This method uses methyl-sensitive restriction enzymes to identify CpG methylation.Refer FIGURE A.
Principle
Methyl-sensitive restriction enzyme cuts only unmethylated DNA and does not cut methylated DNA. If CpG site is methylated, Methyl-sensitive restriction enzyme does not cut DNA And if CpG site is unmethylated, Methyl-sensitive restriction enzyme cuts DNA. The cut DNA fragments are sequenced and counted.
Where fragments are more → methylation is less
Where fragments are less → methylation is more
“That is, number of cut fragments = methylation status.”
Process
i) DNA Extraction – Genomic DNA is isolated from cells
ii) Restriction Enzyme Digestion – DNA is digested with a methyl-sensitive restriction enzyme (e.g. HpaII). This enzyme cuts only at unmethylated CpG sites.
iii) Adapter Ligation – Special adapters are added to the cut DNA fragments.
iv) PCR Amplification – Adapter-ligated fragments are amplified by PCR.
v) High-Throughput Sequencing – Next Generation Sequencing (NGS) of the amplified fragments.
vi) Cut Counting & Data Analysis – Reads from each CpG site are counted (Higher reads → unmethylated and Lower/absent reads → methylated).
Advantage
- High-resolution methylation analysis.
- Genome-wide methylation profiling possible.
- No bisulfite treatment required (DNA damage avoided).
- Quantitative method – accurate quantification of methylation.
- High-throughput sequencing compatible.
- Suitable for CpG-rich regions.
Disadvantage
- Limited to restriction enzyme recognition sites only.
- Does not cover all CpG sites in the entire genome.
- Results depend on enzyme digestion efficiency.
- NGS is expensive.
- Requires complex bioinformatics analysis.
- May give false signals in low-methylated regions.
Application
- Cancer epigenetics studies.
- Developmental biology.
- Gene regulation analysis.
- Comparative methylation studies.
- Finding disease-associated methylation patterns.
B) Affinity-Based Analysis or Affinity-Based Methylation Analysis
It is one of the strategies of Genetic methylation test. Affinity-based Genetic methylation test is a modern method for detecting DNA methylation.In this method, methylated DNA is isolated (enriched) using a special protein or antibody that recognizes methylated DNA. This technique does not require cutting the DNA. Specific binding (affinity) occurs only to DNA regions that have a methyl group (–CH₃) and provides information about methylation.
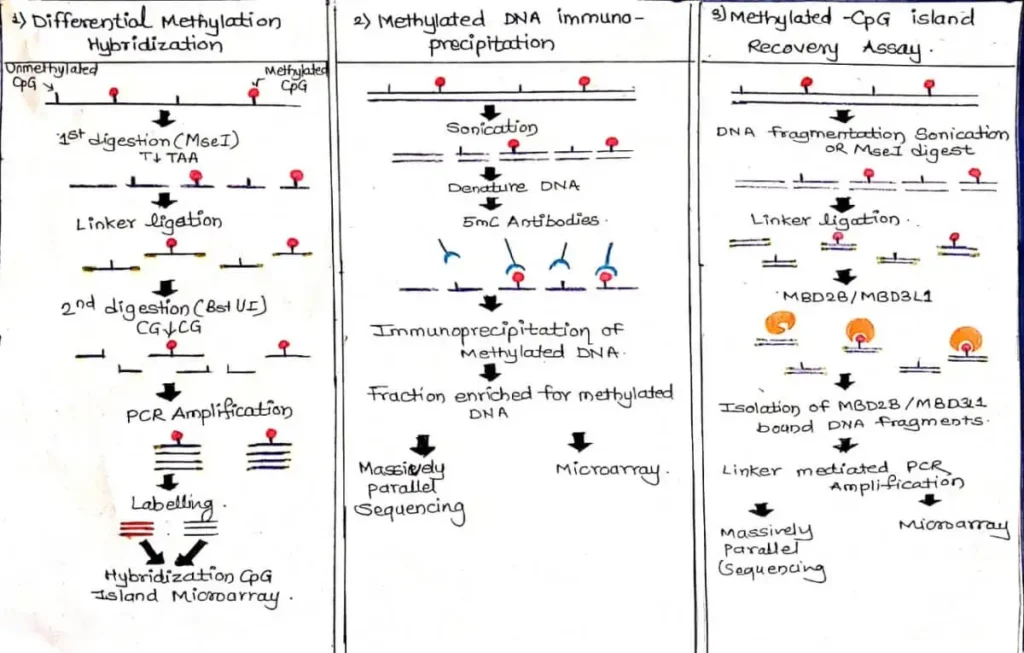
Techniques used in Affinity-Based Methylation Analysis-
1) Differential Methylation Hybridization (DMH)
Differential Methylation Hybridization (DMH) is a microarray-based technique for Genetic methylation test.
This method is used to detect differences in methylation patterns between two samples (e.g. normal vs cancer DNA).Refer FIGURE B.
Principle
In DMH,genomic DNA is fragmented with a methylation-independent restriction enzyme and undergoes adaptor ligation. Next, DNA is digested with the methylation-sensitive enzyme BstUI, PCR amplified, labeled, and cohybridized to CpG island microarrays.
Process
i) DNA Extraction – Genomic DNA is isolated from cells.
ii) First digestion with a methylation-independent restriction enzyme i.e MseI
iii) Adaptor Ligation OR Linker Ligation
iv) Second Digestion with methylation-sensitive enzyme i.e BstUI.
v) PCR Amplification
vi) Labeling – Labeling DNA is labeled with fluorescent dyes (Cy3, Cy5). Normal and test samples are labeled with different colors.
vii) Microarray Hybridization- Labeled DNA is hybridized to a CpG island microarray.
viii) Data Analysis – By comparing signal intensity (Higher signal → higher methylation and Lower signal → lower methylation).The methylation difference between the two samples is detected.
Advantage
- Genome-wide methylation comparison possible.
- Multiple samples can be analyzed together.
- Useful for cancer methylation studies.
- CpG islands can be analyzed well.
- No bisulfite treatment required.
- Relatively fast technique.
Disadvantage
- Limited to regions on array only.
- Single CpG resolution not available.
- Dependent on restriction enzyme sites.
- Low quantitative accuracy.
- High-quality DNA required.
- Less sensitive than NGS.
Appliations
- Cancer vs normal methylation profiling.
- Epigenetic biomarker discovery.
- Gene silencing studies.
- Comparative methylation analysis.
2) Methylated DNA Immunoprecipitation (MeDIP)
MeDIP is an affinity-based Genetic methylation test method. In this technique, methylated DNA is isolated (immunoprecipitated) using a specific antibody that recognizes 5-methylcytosine (5-mC).
In MeDIP, DNA is Split through sonication, denatured, and immunoprecipitated with antibody against 5-methylcytidine. Then Methylated DNA is analyzed using microarrays or sequencing.Refer FIGURE B.
Principle
Antibody specific for 5-methylcytosine binds only to methylated DNA. Methylated cytosine (5-mC) in DNA binds to the antibody. Unmethylated DNA does not bind Antibody–DNA complex is separated and methylation is analyzed. That is, selection of methylated DNA based on antibody affinity.
Process
i) DNA Extraction- Genomic DNA is isolated from cells.
ii) DNA Fragmentation – DNA is broken into smaller fragments by sonication or enzyme.
iii) Denaturation – Double-stranded DNA is made single-stranded(required for antibody binding).
iv) Immunoprecipitation – Anti-5-methylcytosine antibody is added to DNA .Antibody binds only to methylated DNA fragments.
v) Pull Down & Washing – Antibody-DNA complex is separated with beads.Unbound DNA is removed
vi) DNA Recovery – Methylated DNA fragments are recovered.
vii) Analysis – PCR,Microarray (MeDIP-chip),NGS (MeDIP-seq).
Advantage
- Genome-wide methylation analysis possible.
- No restriction enzyme required.
- Useful for CpG-rich regions.
- Suitable for large sample sets.
- Bisulfite treatment can be avoided.
- Widely used in cancer epigenetics.
Disadvantage
- Single CpG resolution not available.
- Results depend on antibody specificity.
- Difficult to identify low-density methylation.
- Low quantitative accuracy.
- Bias may occur in GC-rich regions.
- Absolute methylation percentage not available.
Applications
- Cancer methylation profiling.
- Search Epigenetic biomarker .
- Gene silencing studies.
- Comparative methylation analysis.
- Developmental epigenetics.
3) Methyalated-CpG island Recovery Assay (MIRA)
MIRA is an Affinity-based Genetic methylation test method.In this technique, methylated CpG-rich DNA (CpG islands) are specifically isolated (enriched) using MBD (Methyl-CpG Binding Domain) proteins.
In MIRA, DNA split by sonication or MseI digestion undergoes adaptor ligation followed by incubation with MBD2b/MBD3L1 proteins. The MIRA captured DNA is then PCR amplified and analyzed using microarrays or sequencing.Refer FIGURE B.
Principle
MBD proteins bind specifically to methylated CpG sequences only.(Methylated CpG islands → bind to MBD proteins and Unmethylated CpG islands → do not bind).The bound DNA is isolated and methylation is analyzed .That is, methylated CpG islands are identified based on protein affinity.
Process
i) DNA Extraction – Genomic DNA is isolated from cells.
ii) DNA Fragmentation – DNA is broken into small fragments by sonication or enzyme.
iii) Binding with MBD Proteins- DNA fragments are incubated with the MBD2b and MBD3L1 protein complex.This complex binds only to methylated CpG-rich DNA.
iv) Pull-Down of Methylated DNA -The protein–DNA complex is separated using beads.
v) Washing – Washing is performed to remove unmethylated DNA.
vi) DNA Recovery – Methylated CpG island DNA is recovered.
vii) Analysis – Analysis of DNA obtained:PCR,Microarray (MIRA-chip,NGS (MIRA-seq).
Advantage
- Highly specific for CpG islands.
- No antibody required (more stable than MeDIP).
- High signal-to-noise ratio.
- Genome-wide methylation profiling possible.
- Low background binding.
- Useful for cancer methylation studies.
Disadvantage
- Non-CpG methylation is not detected.
- Single CpG resolution is not available.
- Not suitable for CpG-poor regions.
- Quantitative methylation percentage is not available.
- Depends on protein binding efficiency.
- Absolute methylation level cannot be determined.
Applications
- CpG island methylation profiling.
- Finding cancer-specific methylation patterns.
- Gene promoter methylation studies.
- Epigenetic biomarker identification.
C) Bisulfite treatment OR Bisulfite-Based Analysis
It is one of the strategies of Genetic methylation test. Bisulfite treatment is a chemical process used to identify methylation in DNA. In this process, DNA is treated with sodium bisulfite. In this, unmethylated cytosine is converted to uracil, while methylated cytosine does not undergo any change. Therefore, the difference between methylated and unmethylated DNA can be clearly identified in subsequent PCR or sequencing methods.This method is extremely important in Genetic methylation test and epigenetic studies.
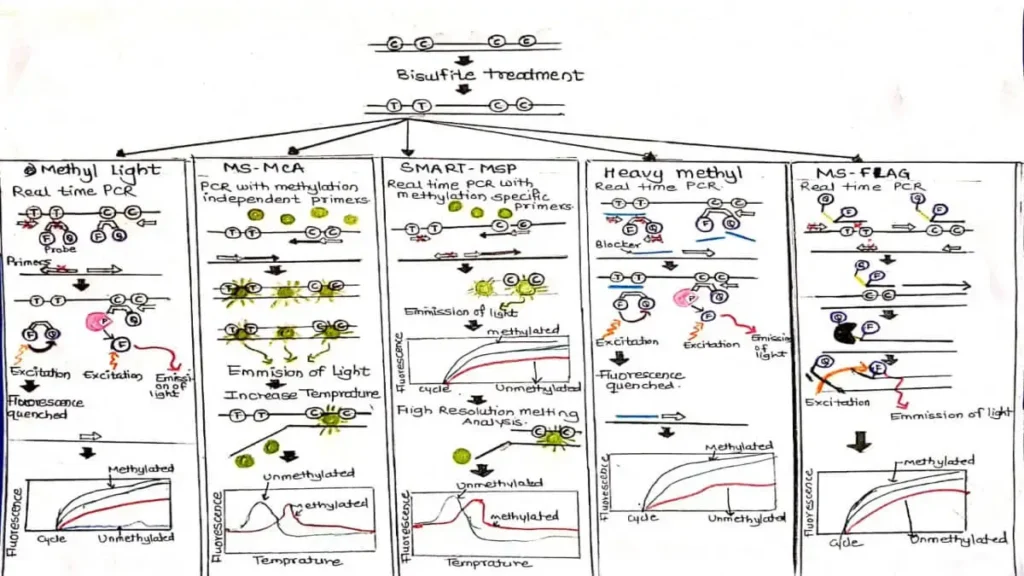
1) Methyl Light-
Principle
DNA is first given Sodium Bisulfite treatment. In this treatment Unmethylated cytosine → changes to uracil and Methylated cytosine does not. Then Real-Time PCR (qPCR) is performed using methylation-specific primers and probes.Only methylated DNA sequences are amplified.The amount of methylation is measured based on the fluorescence signal.REFER FIGURE C 1.
Process
i) DNA Extraction – Genomic DNA is isolated from cells or tissue.
ii) Bisulfite Treatment – DNA is treated with sodium bisulfite. unmethylated cytosine converted to uracil.
iii) Primer & Probe Design – Primers and probes are designed that recognize only methylated DNA.
iv) Real-Time PCR (qPCR) – A fluorescence signal is generated during PCR. The intensity of the signal is used to measure methylation.
v) Data Analysis – The methylation level is quantified using the Ct value.
Advantage
- Highly sensitive method.
- Quantitative analysis possible (measures methylation).
- Works even with low amounts of DNA.
- Fast and high-throughput technique.
- Useful for identifying cancer biomarkers.
- Suitable for automation.
Disadvantage
- Can only be used for known gene regions.
- Primer design is very important and difficult.
- Bisulfite treatment can degrade DNA.
- Requires expensive equipment (Real-time PCR machine).
- Genome-wide methylation analysis is not possible.
2) Methylation Sensitive Melting Curve Analysis(MS-MCA)
MS-MCA is a Real-Time PCR based method used to detect methylation at CpG sites in DNA. This method is based on the difference in DNA melting temperature (Tm).
DNA is first given bisulfite treatment. After this treatment, Unmethylated cytosine → changes to uracil (then thymine) and Methylated cytosine does not change.Therefore, the DNA fragments produced after PCR have different GC content. Due to the difference in GC content,The melting temperature (Tm) of methylated and unmethylated DNA is different. This difference is identified by melting curve analysis..REFER FIGUREC1
Process
i) DNA Extraction – Genomic DNA is isolated from cells or tissue.
ii) Bisulfite Treatment – DNA is treated with sodium bisulfite.Unmethylated cytosine convert in urail and then thymine.
iii) PCR Amplification – PCR is performed using methylation-independent primers. The reaction is carried out in a real-time PCR machine.
iv) Melting Curve Analysis – After PCR, the temperature is gradually increased. DNA double strand convered in single strand and Fluorescence decreases.
v) Data Interpretation – By observing the melting peaks (Tm),methylated and unmethylated or mixed methylation pattern is identified.
Advantage
- Simple and rapid technique.
- Quantitative prediction is possible.
- Closed-tube system reduces contamination.
- No need for gel electrophoresis.
- Partially methylated DNA can be detected.
- Cost-effective (cheaper than MethyLight).
Disadvantage
- Primer design needs to be done very carefully.
- Interpretation becomes difficult if Tm difference is low.
- High-resolution instrument required..
- Genome-wide methylation analysis is not possible.
- Bisulfite treatment can damage DNA.
3) Sensitive Melting Analysis After Real-Time (SMART-MSP)
Priniple
In SMART-MSP, bisulfite-modified DNA undergoes methylation-specific amplification in the
presence of double-stranded intercalating dye such as SYBR green (represented by green circles) and the amount of signal detected is proportional to the amount of methylated DNA. Following PCR, the reaction temperature is increased and DNA melting properties are examined..REFER FIGUREC1.
Advantage
- Very high sensitivity – Even very low levels of methylated DNA can be detected.
- Real-time PCR + Melting analysis combined -Both amplification and analysis in a single reaction.
- Partially methylated DNA detection possible – Mixed methylation pattern can be identified.
- No need for gel electrophoresis – Saves time and effort.
- Closed-tube system – Reduced risk of contamination.
- Useful for cancer biomarker studies – Useful for identifying early stage methylation.
Disadvantage
- Primer design is very critical – Wrong primer can give false results.
- High-resolution real-time PCR machine required – Cost is high.
- Interpretation difficult if Tm difference is low – Methylation levels may not be clearly differentiated.
- Bisulfite treatment required – DNA degradation possible.
- Genome-wide methylation analysis not possible – Limited to specific gene regions only.
4) Heavy Methyl
Principle and Process
HeavyMethyl utilizes blocker oligonucleotides that specifically bind to unmethylated DNA and prevent its amplification. Alternatively,methylated DNA is amplified using methylation-independent primers and a methylation-specific probe that contains a fluorophore (F) and a quencher (Q). During the PCR reaction, the probe is cleaved by the exonuclease activity of DNA polymerase, causing the fluorophore to be released from the quencher and light to be emitted. The emitted light signal is proportional to the amount of methylated DNA present in the sample.REFER FIGURE C.1.
Advantage
HeavyMethyl is a highly sensitive and specific method that can accurately detect very low levels of methylated DNA even in the presence of background unmethylated DNA.
Disadvantage
This technique is costly and complicated because it requires bisulfite treatment, blocking oligonucleotides, expensive reagents, and complex primer design.
5) Methylation-Specific Fluorescent Amplicon Generation(MS-FLAG)
Principle and Process
In MS-FLAG, bisulfite-treated DNA is amplified with methylation-specific primers that contain a cleavage site for PspGI. Additionally, the primers contain a fluorophore (F) and a quencher (Q). The cleavage of the primers by PspGI enables the release of the quencher from the fluorophore and light to be emitted, which is proportional to amount of methylated DNA.REFER FIGURE C.1.
Advantage
Methylation-Specific Fluorescent Amplicon Generation (MS-FLAG) method offers high specificity and sensitivity and accurately detects methylated DNA through fluorescent signal, thus eliminating the need for gel electrophoresis.
Disadvantage
This technique requires bisulfite treatment, special fluorescent probes/primers, and expensive equipment, making it expensive and not capable of genome-wide methylation analysis.
6) Epi TYPER
Principle and Process
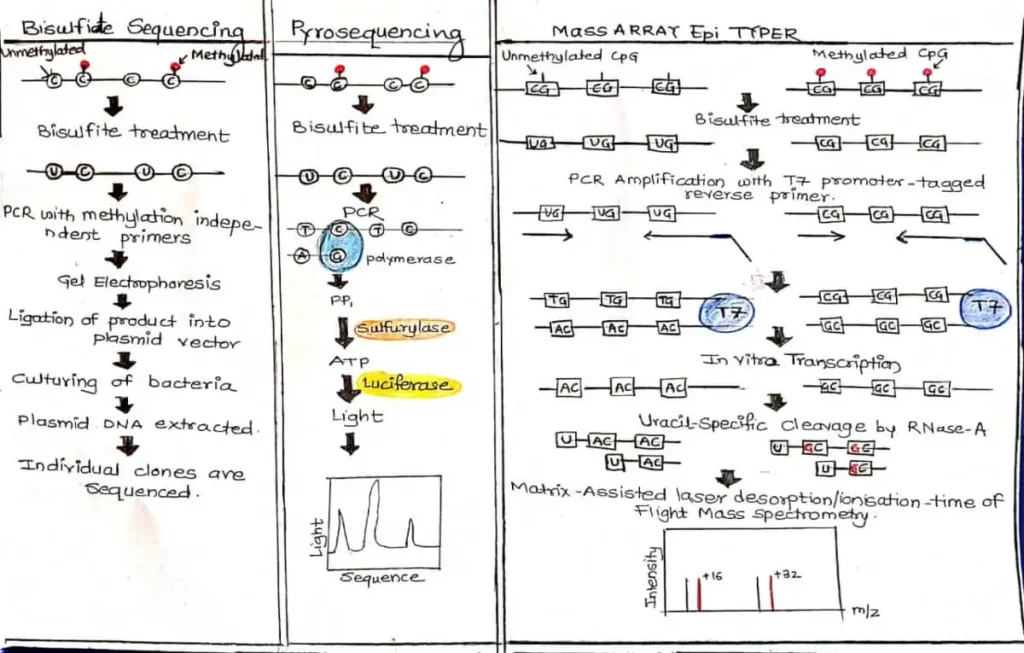
PCR amplification of Bisulfite-modified DNA with T7 Promotor-taggedreverse primer. Then, in vitro RNA transcription is performed and then digestion with RNase .After that Analysis of digestion products by MALDI-TOF MS . Methylated cytosines are transcribed to guanine, whereas unmethylated cytosines are converted to uracils and transcribed to adenines. This is represented in the mass spectrum by signal pairs separate by 16 m/z (or multiples thereof).FIGURE C.2.
Advantage
The EpiTYPER method provides CpG site-specific and quantitative methylation analysis, allowing the precise methylation pattern of multiple CpG sites to be identified simultaneously.
Disadvantage
This technique requires MALDI-TOF mass spectrometry, bisulfite treatment, and specialized software, making it expensive and the process technically complex.
7) Bisulfite Sequencing
Principle and Process
In bisulfite sequencing,PCR amplification of bisulfite-treated DNA is with methylation-independent primers and size fractioned using gel electrophoresis. The purified PCR products are then cloned intoE. coli and individual clones (usually 5–10) are sequenced.FIGURE C.2.
Advantage
Bisulfite sequencing is the gold-standard method for Genetic methylation analysis and provides highly accurate and quantitative information at single-base (CpG site-level) resolution.
Disadvantage
This technique is rarely used for routine screening due to the possibility of DNA damage due to bisulfite treatment, as well as the high cost, time, and data analysis complexity.
8) Pyrosequencing
Principle and Process
In pyrosequencing, bisulfite-modified DNA is amplified with DNA polymerase and sequencing primers. As the complementary DNA strand is synthesized, PPi is released and converted into ATP. The ATP provides the energy to form a luciferase–luciferin–AMP complex, which in the presence of oxygen results in the release of light in a proportional amount to the available ATP and thus PPi.FIGURE C.2.
Advantage
Pyrosequencing provides CpG site–specific and quantitative methylation analysis, resulting in accurate, reproducible, and real-time sequencing results.
Disadvantage
This technique requires bisulfite treatment, special reagents, and equipment, making it expensive and limiting for long DNA sequences.
Conclusion
Genetic methylation testing is considered a revolutionary and highly reliable molecular method in the field of epigenetics today.Genetic Methylation test provides in-depth and accurate information about how gene expression is controlled, how diseases begin, and what molecular changes occur early in diseases like cancer. In Genetic Methylation test ,Modern bisulfite-based, PCR-based, and sequencing-based techniques provide highly sensitive, quantitative, and reproducible methylation data at the CpG site-level.
When the appropriate Genetic Methylation test method is chosen considering the purpose of the study, sample quality, desired resolution, and cost, genetic methylation testing can be a powerful and future-oriented scientific tool for early disease detection, biomarker identification, and personalized medicine.\
The genetic methylation test is expected to play an increasingly important role in the future of genomics, epigenetics, and precision medicine.In the coming years, genetic methylation testing is likely to support early disease prediction and risk stratification, especially for complex conditions such as cancer, cardiovascular disorders, neurological diseases, and metabolic syndromes.
As scientific understanding of DNA methylation deepens, genetic methylation test will move beyond research and wellness applications toward mainstream clinical and preventive healthcare
References
1. Waddington, C. H. 1942. The epigenotype. Endeavour 1:18–20.
2.Ekaterina Olkhov-Mitse & Bharati Bapat 2012. Strategies for discovery and validation of methylated and
hydroxymethylated DNA biomarkers.Cancer Medicine 2012; 1(2): 237–260.
3.Haedicke, W., and R. Lesche. 2006. DNA methylation:potential for diagnosis, prognosis and therapy – prediction in oncology. Verh. Dtsch. Ges. Pathol. 90:39–45.
4.Sergey Kurdyukov, Martyn Bullock .Biology (Basel). 2016 Jan 6;5(1):3.
Also Read
15+ Important Scope of Microbiology – Best Jobs After Microbiology
What is a genetic methylation test?
methylation test is a DNA-based test that analyzes how well your body performs DNA methylation—a key biochemical process that regulates gene expression, detoxification, neurotransmitter balance, hormone metabolism, and cardiovascular health. This test usually examines specific gene variants (SNPs) involved in the methylation cycle, such as MTHFR, MTR, MTRR, COMT, CBS, and BHMT, to understand how efficiently your body uses nutrients like folate, vitamin B12, B6, and choline.In simple terms, it helps explain how your genes process nutrients and control important metabolic pathways.
What does a genetic methylation test tell you?
A genetic methylation test can provide insights into:
How efficiently your body performs methylation reactions.
Your ability to convert folate into its active form.
Risk of elevated homocysteine levels.
How your body handles detoxification and oxidative stress.
Neurotransmitter balance (dopamine, serotonin, adrenaline).
Why certain vitamins or supplements work—or don’t work—for you.
Personalized nutrition and lifestyle optimization clue.
Where to get a genetic methylation test?
You can get a genetic methylation test from:
1.Direct-to-consumer genetic testing companies.
2.Functional medicine clinics.
3.Preventive health and genomics labs.
4.Some hospitals or diagnostic centers (via doctors).
Where can I get a genetic methylation test?
You can obtain the test through:
1.Online genetic testing platforms (home test kits).
2.Functional medicine practitioners.
3.Integrative or lifestyle medicine doctors.
4.Wellness and biohacking clinics.
After sample submission, results are typically available within 2–4 weeks, often with a downloadable report.
How much is a genetic methylation test?
The cost of a genetic methylation test generally ranges between:
Basic test: ₹8,000 – ₹15,000 (or $100–$200)
Advanced panels with interpretation: ₹20,000 – ₹40,000 (or $250–$500)
Pricing depends on:
1.Number of genes analyzed.
2.Whether expert interpretation is included.
3.Lab technology used.
What is genetic methylation test used for?
Genetic methylation testing is commonly used for:
1.Preventive and personalized health planning.
2.Nutritional genomics.
3.Functional medicine assessments.
4.Understanding fatigue, brain fog, mood issues, or nutrient deficiencies.
5.Supporting lifestyle and supplement decisions.